 In the blog post Configuring a Neural Network Output Layer we highlighted how to correctly set up an output layer for deep learning models. Here, we discuss how to make sense of what a neural network actually returns from the output layers. If you are like me, you may have been surprised when you first encountered the output of a simple classification neural net.
In the blog post Configuring a Neural Network Output Layer we highlighted how to correctly set up an output layer for deep learning models. Here, we discuss how to make sense of what a neural network actually returns from the output layers. If you are like me, you may have been surprised when you first encountered the output of a simple classification neural net.
“Wait… I thought this was supposed to give me an array of integers that was similar to my target variable. Why am I seeing floating point numbers? What do these numbers even mean?”
In this post, we’ll answer these types of questions and more by showing how to extract target labels from deep learning classification models.
Classification Review
A typical classification problem differs from a regression problem in that we are trying to predict labels or classes as a target rather than some continuous value. In a regression problem, the output layer needs no activation function – a simple linear transform can be applied to the result of the neural network and this output can be directly interpreted in the context of the target variable. With classification, this is not the case; either a sigmoid or softmax wrapper is needed to convert the output of the model into probability distributions. The classification problem type binary, multi-class, or multi-label determines how we interpret these results. If you need a refresher on this, review our previous post here before continuing on.
In order to compare the various classification problem types, we will set up a simple example problem. Let’s assume we are tasked with detecting whether or not any of the noble gasses are present in an environment based on a collection of observed features.
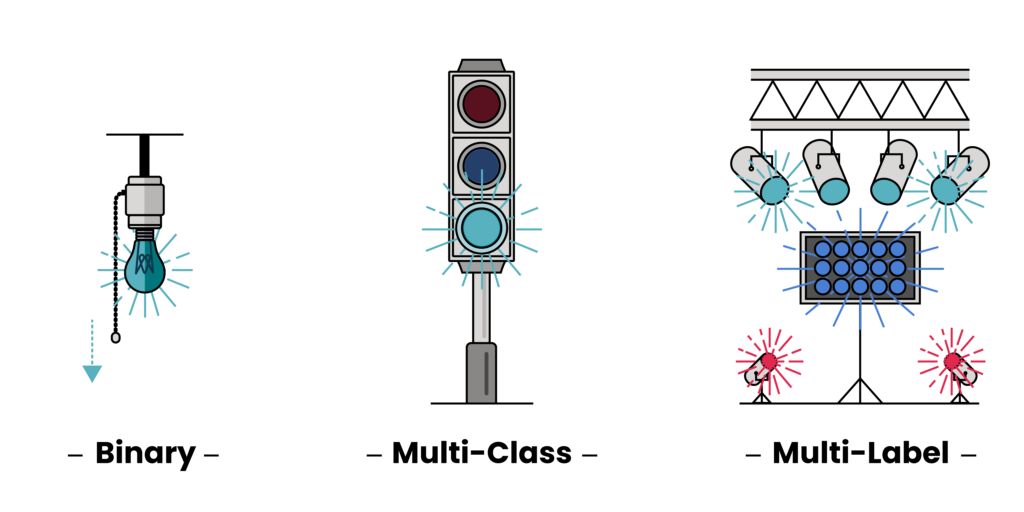
For a binary classification problem, this would be determining if any of the gasses were present. This would be like asking: “Given this collection of features, do I believe any of the noble gasses—Helium (He), Neon (Ne), Argon (Ar), Krypton (Kr), Xenon (Xe), or Radon (Rn)—exist in this environment?” For a multi-class problem, we’d have to refine our search and look for the individual most concentrated noble gas. This would be like asking: “Given this collection of features, which single noble gas do I think is in this environment?” Finally, for a multi-label problem, we’d update our search to look for any and all noble gasses: “Given this collection of features, which combination of the noble gasses is in this environment?”
Why Extract Labels?
Extracting target labels is not a necessary task for a deep learning model to perform. From a model’s perspective, keeping things in an integer or floating point state is often required. However, to us humans, converting things from numeric form to textual form provides great value. For example, consider the final predictions of a classification model to be the array [0, 1, 0, 2, 0, ...]. Furthermore, consider the confusion matrix between these classes to be the below matrix:
[[50, 0, 0],
[ 0, 47, 3],
[ 0, 1, 49]]
These numbers don’t mean much to us without the problem context. If I were to tell you that the class labels were ['red', 'green', 'blue'], now you’d be able to interpret this output more efficiently. You’d now know that the first 5 predictions from the above array were ‘red‘, ‘green‘, ‘red‘, ‘blue‘ and ‘red‘ respectively. You’d also be able to see that all the ‘red‘ samples were correctly labeled while the ‘green‘ and ‘blue‘ classes were harder to distinguish (3 true ‘green‘ mislabeled as ‘blue‘; 1 true ‘blue‘ mislabeled as ‘green‘). As shown here, extracting target labels from a model can be extremely valuable to us humans and can help us interpret predictions and better understand our model’s performance.
Extraction Through Example
The different noble gas questions listed above give us context to describe how each of the classification problem types treat the output probability distribution from a deep learning model.
Binary Classification
In binary classification, a sigmoid activation function is used to convert model output to probability distributions relating to only one class. Here, each output probability must be compared to a threshold value to determine whether or not the prediction belongs to class 1.
>>> import tensorflow as tf
>>> tf.random.set_seed(42)
# Random prediction output for 5 samples
# logits means the vector of raw (non-normalized) predictions
>>> logits = tf.random.normal([5,], mean=0, stddev=5, seed=13)
>>> print(logits.numpy())
[ 5.849211 -0.15770258 0.04067278 -5.5301137 2.3940122 ]
# Mimic output layer of neural network for 5 samples
>>> binary_out = tf.nn.sigmoid(logits)
>>> print(binary_out.numpy())
[0.9971261 0.46065587 0.51016676 0.00394991 0.91636956]
# Interpret probability distribution with 0.5 threshold
>>> binary_preds = tf.cast(binary_out >= 0.5, tf.int32)
>>> print(binary_preds.numpy())
[1 0 1 0 1]
In the above binary case, we use 0.5 as the threshold to determine which binary class the output prediction belongs to: >= 0.5 means class 1 and < 0.5 means not class 1 (i.e., class 0). If we recall our binary question, “Given this collection of features, do I believe any of the noble gasses exist in this environment,” we could say that our target labels pertain to detected and nothing-detected class labels. Here, extracting target labels is straightforward—we can perform a lookup based on index to get the corresponding prediction labels.
>>> import numpy as np
>>> binary_target_names = np.array([‘nothing-detected’, ‘detection’])
>>> binary_preds_labeled = binary_target_names[binary_preds.numpy()]
>> print(binary_preds.numpy())
[1 0 1 0 1]
>>> print(binary_preds_labeled)
[‘detection’ ‘nothing-detected’ ‘detection’ ‘nothing-detected’ ‘detection’]
Multi-Class Classification
Now, let’s shift focus to the multi-class problem type. Below, we create another mock output for 5 samples, but this time, instead of a single value, we get an output for each of the target classes in our problem. Here, that means we have a floating point value for each of the noble gasses: Helium (He), Neon (Ne), Argon (Ar), Krypton (Kr), Xenon (Xe), and Radon (Rn). We then use the softmax activation function to convert to probability distributions.
# Random prediction output for 5 samples
>>> logits = tf.random.normal([5,6], mean=0, stddev=5, seed=42)
# Set numpy print options to only show two decimal places for display
>>> np.set_printoptions(precision=2)
>>> print(logits.numpy())
[[ 6.57 -0.77 4.56 -4. -0.54 1.42]
[ 3.83 -3.11 4.99 1.1 -5.4 -9.94]
[ 7.8 -0.85 -6.89 -1.49 4.21 4.61]
[-2.42 -8.49 0.48 -0.53 -8.92 8.57]
[-5.88 5.74 2.25 -4.09 4.76 6.96]]
# Mimic output layer of neural network for 5 samples
>>> multi_class_out = tf.nn.softmax(logits)
>>> print(multi_class_out.numpy())
[[8.77e-01 5.66e-04 1.17e-01 2.25e-05 7.11e-04 5.07e-03]
[2.36e-01 2.29e-04 7.49e-01 1.53e-02 2.31e-05 2.47e-07]
[9.36e-01 1.64e-04 3.89e-07 8.62e-05 2.56e-02 3.86e-02]
[1.68e-05 3.90e-08 3.05e-04 1.11e-04 2.53e-08 1.00e+00]
[1.88e-06 2.10e-01 6.39e-03 1.13e-05 7.84e-02 7.05e-01]]
For a multi-class problem, we have multiple target labels but our prediction must be mutually exclusive. Recall the question for our example: “Given this collection of features, which single noble gas do I think is in this environment?” Using the above multi-class output, we can use the argmax method to extract the noble gas with the highest probability of detection.
# Interpret probability distribution with
# highest probability as chosen class
>>> multi_class_preds = multi_class_out.numpy().argmax(axis=1)
>>> print(multi_class_preds)
[0 2 0 5 5]
>>> noble_gasses = np.array(['He', 'Ne', 'Ar', 'Kr', 'Xe', 'Rn'])
>>> multi_class_preds_labeled = noble_gasses[multi_class_preds]
>>> print(multi_class_preds_labeled)
['He' 'Ar' 'He' 'Rn' 'Rn']
Multi-Label Classification
When moving to multi-label classification, things get a bit more complicated. In this type of classification problem, we are trying to determine any number of labels in which we don’t have to satisfy mutual exclusivity. Our question now becomes, which collection of noble gasses are being detected rather than which single gas will be predicted.
As before, we start by creating our logits array—a mock vector of raw (non-normalized) predictions that a classification model generates—for 5 samples. Then, we pass this to the sigmoid activation function to generate our probability distributions for each class.
# Random prediction output for 5 samples
>>> logits = tf.random.normal([5,6], mean=0, stddev=5, seed=13)
>>> print(logits.numpy())
[[-8.71 -0.2 0.12 0.78 -1.37 -2.86]
[ 1.27 2.8 -0.17 -1.92 12.42 6.13]
[-4.49 -1.74 -2.68 -6.44 -3.47 -7.65]
[ 5.09 6.38 8.65 1.05 -9.85 6.11]
[ 2.41 0.12 0.27 -9.25 3.36 2.2 ]]
# Mimic output layer of neural network for 5 samples
>>> multi_label_out = tf.nn.sigmoid(logits)
>>> print(multi_label_out.numpy())
[[1.64e-04 4.49e-01 5.30e-01 6.87e-01 2.02e-01 5.43e-02]
[7.81e-01 9.43e-01 4.57e-01 1.28e-01 1.00e+00 9.98e-01]
[1.11e-02 1.49e-01 6.41e-02 1.60e-03 3.02e-02 4.74e-04]
[9.94e-01 9.98e-01 1.00e+00 7.41e-01 5.30e-05 9.98e-01]
[9.17e-01 5.29e-01 5.67e-01 9.59e-05 9.66e-01 9.00e-01]]
When we are presented a multi-label problem, we need to fit multiple binary classifications for each target class. By classifying probabilities that have a value greater than or equal to 0.5 as 1, and those values less than 0.5 as 0, we can detect multiple classes in our outputs. As shown below, any row that has at least one 1 indicates that a noble gas was detected. The column index of the 1 corresponds to the target labels, noble_gasses, defined in the multi-class section above. The question here is, “Given this collection of features, which combination of the noble gasses are in this environment?”
# Interpret probability distribution with 0.5 threshold
>>> multi_label_preds = tf.cast(
... multi_label_out.numpy() >= 0.5, tf.int32
... )
>>> print(multi_label_preds.numpy())
[[0 0 1 1 0 0]
[1 1 0 0 1 1]
[0 0 0 0 0 0]
[1 1 1 1 0 1]
[1 1 1 0 1 1]]
The multi-label problem type introduces a unique label extraction problem: we can’t simply look up class labels based on an index.
>>> for row in multi_label_preds.numpy():
... print(noble_gasses[row])
['He' 'He' 'Ne' 'Ne' 'He' 'He']
['Ne' 'Ne' 'He' 'He' 'Ne' 'Ne']
['He' 'He' 'He' 'He' 'He' 'He']
['Ne' 'Ne' 'Ne' 'Ne' 'He' 'Ne']
['Ne' 'Ne' 'Ne' 'He' 'Ne' 'Ne']
Hmm, that’s not right. What if we select based on the non-zero values?
>>> for row in multi_label_preds.numpy():
... print(noble_gasses[row.nonzero()])
['Ar' 'Kr']
['He' 'Ne' 'Xe' 'Rn']
[]
['He' 'Ne' 'Ar' 'Kr' 'Rn']
['He' 'Ne' 'Ar' 'Xe' 'Rn']
That’s better!
Conclusion
Classification is one of the most popular problems encountered in deep learning. In this post, we’ve taken a closer look at how to extract target labels from various classification problem types. For binary cases, we showed how to use a simple indexing approach to move from 1s and 0s to more meaningful target label descriptors. For multi-class problems, we discussed the use of numpy.argmax() to select the highest probability from the output distribution and map it to a class label. Finally, we discussed the multi-label problem and showed how to use the numpy.nonzero() method to extract labels from each sample prediction where mutually exclusivity is not required. With these new skills, you’re ready to extract labels from all types of classification problems and convert numerical outputs to meaningful class labels.
Happy extracting!
About the Author
Logan Thomas holds a M.S. in mechanical engineering with a minor in statistics from the University of Florida and a B.S. in mathematics from Palm Beach Atlantic University. Logan has worked as a data scientist and machine learning engineer in both the digital media and protective engineering industries. His experience includes discovering relationships in large datasets, synthesizing data to influence decision making, and creating/deploying machine learning models.
Related Content
Digital Transformation vs. Digital Enhancement: A Starting Decision Framework for Technology Initiatives in R&D
Leveraging advanced technology like generative AI through digital transformation (not digital enhancement) is how to get the biggest returns in scientific R&D.
Digital Transformation in Practice
There is much more to digital transformation than technology, and a holistic strategy is crucial for the journey.
Leveraging AI for More Efficient Research in BioPharma
In the rapidly-evolving landscape of drug discovery and development, traditional approaches to R&D in biopharma are no longer sufficient. Artificial intelligence (AI) continues to be a...
Utilizing LLMs Today in Industrial Materials and Chemical R&D
Leveraging large language models (LLMs) in materials science and chemical R&D isn't just a speculative venture for some AI future. There are two primary use...
Top 10 AI Concepts Every Scientific R&D Leader Should Know
R&D leaders and scientists need a working understanding of key AI concepts so they can more effectively develop future-forward data strategies and lead the charge...
Why A Data Fabric is Essential for Modern R&D
Scattered and siloed data is one of the top challenges slowing down scientific discovery and innovation today. What every R&D organization needs is a data...
Jupyter AI Magics Are Not ✨Magic✨
It doesn’t take ✨magic✨ to integrate ChatGPT into your Jupyter workflow. Integrating ChatGPT into your Jupyter workflow doesn’t have to be magic. New tools are…
Top 5 Takeaways from the American Chemical Society (ACS) 2023 Fall Meeting: R&D Data, Generative AI and More
By Mike Heiber, Ph.D., Materials Informatics Manager Enthought, Materials Science Solutions The American Chemical Society (ACS) is a premier scientific organization with members all over…
Real Scientists Make Their Own Tools
There’s a long history of scientists who built new tools to enable their discoveries. Tycho Brahe built a quadrant that allowed him to observe the…
How IT Contributes to Successful Science
With the increasing importance of AI and machine learning in science and engineering, it is critical that the leadership of R&D and IT groups at...